Abstract
β-Thalassaemia is caused by reduced or absent production of β-haemoglobin1,2,3,4. Previously, we performed laboratory-scale electroporation of CD34+ haematopoietic stem and progenitor cells from patients with β-thalassaemia using a transformer base editor5,6. The aim was to target the binding motif of the transcription repressor BCL11A in the HBG1 and HBG2 promoters7 to reactivate fetal haemoglobin (HbF) production. Here we present results of a phase 1 clinical trial (ClinicalTrials.gov identifier: NCT06024876) of five patients who received autologous CD34+ cells modified using a transformer base editor at clinical scale (CS-101). With a median follow-up of 23.0 months after CS-101 infusion, the median times to neutrophil and platelet engraftment were 16 days and 25 days, respectively. Moreover, all patients had stopped red blood cell transfusions, with a median time to the last transfusion of 18 days after CS-101 infusion. The mean total haemoglobin and HbF concentrations were 12.4 ± 1.0 and 11.5 ± 0.9 g dl–1, respectively, at month 3 after infusion. These levels remained at similar or higher levels throughout the follow-up period, which indicated rapid haematopoietic reconstitution. The adverse events of CS-101 were generally consistent with those of busulfan myeloablative conditioning and autologous haematopoietic stem and progenitor cell transplantation. No deaths or cancer occurrences were reported. In summary, CS-101 can lead to rapid and sustained increases in both total haemoglobin and HbF levels, which resulted in early and enduring transfusion independence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The original DNA sequencing data from this study are currently being deposited in the Sequence Read Archive (SRA) database (accession number: PRJNA1222469). Raw gel images are included as Supplementary Fig. 1. Source data are provided with this paper.
Code availability
The custom Perl and Shell scripts for calculating frequencies of base substitution are available at GitHub (https://github.com/YangLab/CFBI).
References
Cao, A. & Galanello, R. Beta-thalassemia. Genet. Med. 12, 61–76 (2010).
Origa, R. Beta-thalassemia. Genet. Med. 19, 609–619 (2017).
Taher, A. T., Musallam, K. M. & Cappellini, M. D. β-Thalassemias. N. Engl. J. Med. 384, 727–743 (2021).
Hardouin, G., Miccio, A. & Brusson, M. Gene therapy for β-thalassemia: current and future options. Trends Mol. Med. https://doi.org/10.1016/j.molmed.2024.12.001 (2025).
Wang, L. et al. Eliminating base-editor-induced genome-wide and transcriptome-wide off-target mutations. Nat. Cell Biol. 23, 552–563 (2021).
Han, W. et al. Design and application of the transformer base editor in mammalian cells and mice. Nat. Protoc. 18, 3194–3228 (2023).
Han, W. et al. Base editing of the HBG promoter induces potent fetal hemoglobin expression with no detectable off-target mutations in human HSCs. Cell Stem Cell 30, 1624–1639 (2023).
Galanello, R. & Origa, R. Beta-thalassemia. Orphanet J. Rare Dis. 5, 11 (2010).
Thein, S. L. The molecular basis of β-thalassemia. Cold Spring Harb. Perspect. Med. 3, a011700 (2013).
Cavazzana-Calvo, M. et al. Transfusion independence and HMGA2 activation after gene therapy of human β-thalassaemia. Nature 467, 318–322 (2010).
Thompson, A. A. et al. Gene therapy in patients with transfusion-dependent β-thalassemia. N. Engl. J. Med. 378, 1479–1493 (2018).
Locatelli, F. et al. Betibeglogene autotemcel gene therapy for non-β0/β0 genotype β-thalassemia. N. Engl. J. Med. 386, 415–427 (2022).
Li, S. et al. Modified lentiviral globin gene therapy for pediatric β0/β0 transfusion-dependent β-thalassemia: a single-center, single-arm pilot trial. Cell Stem Cell 31, 961–973 (2024).
Kwiatkowski, J. L. et al. Betibeglogene autotemcel gene therapy in patients with transfusion-dependent, severe genotype β-thalassaemia (HGB-212): a non-randomised, multicentre, single-arm, open-label, single-dose, phase 3 trial. Lancet 404, 2175–2186 (2024).
Bauer, D. E. et al. An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science 342, 253–257 (2013).
Canver, M. C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015).
Traxler, E. A. et al. A genome-editing strategy to treat β-hemoglobinopathies that recapitulates a mutation associated with a benign genetic condition. Nat. Med. 22, 987–990 (2016).
Musallam, K. M. et al. Fetal hemoglobin levels and morbidity in untransfused patients with β-thalassemia intermedia. Blood 119, 364–367 (2012).
Bauer, D. E. & Orkin, S. H. Hemoglobin switching’s surprise: the versatile transcription factor BCL11A is a master repressor of fetal hemoglobin. Curr. Opin. Genet. Dev. 33, 62–70 (2015).
Canver, M. C. & Orkin, S. H. Customizing the genome as therapy for the β-hemoglobinopathies. Blood 127, 2536–2545 (2016).
Wienert, B., Martyn, G. E., Funnell, A. P. W., Quinlan, K. G. R. & Crossley, M. Wake-up sleepy gene: reactivating fetal globin for β-hemoglobinopathies. Trends Genet. 34, 927–940 (2018).
Sankaran, V. G. et al. Human fetal hemoglobin expression is regulated by the developmental stage-specific repressor BCL11A. Science 322, 1839–1842 (2008).
Wu, Y. et al. Highly efficient therapeutic gene editing of human hematopoietic stem cells. Nat. Med. 25, 776–783 (2019).
Zeng, J. et al. Therapeutic base editing of human hematopoietic stem cells. Nat. Med. 26, 535–541 (2020).
Frangoul, H. et al. CRISPR–Cas9 gene editing for sickle cell disease and β-thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Fu, B. et al. CRISPR–Cas9-mediated gene editing of the BCL11A enhancer for pediatric β0/β0 transfusion-dependent β-thalassemia. Nat. Med. 28, 1573–1580 (2022).
Locatelli, F. et al. Exagamglogene autotemcel for transfusion-dependent β-thalassemia. N. Engl. J. Med. 390, 1663–1676 (2024).
Haapaniemi, E., Botla, S., Persson, J., Schmierer, B. & Taipale, J. CRISPR–Cas9 genome editing induces a p53-mediated DNA damage response. Nat. Med. 24, 927–930 (2018).
Schiroli, G. et al. Precise gene editing preserves hematopoietic stem cell function following transient p53-mediated DNA damage response. Cell Stem Cell 24, 551–565 (2019).
Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat. Biotechnol. 36, 765–771 (2018).
Cullot, G. et al. CRISPR–Cas9 genome editing induces megabase-scale chromosomal truncations. Nat. Commun. 10, 1136 (2019).
Tsuchida, C. A. et al. Mitigation of chromosome loss in clinical CRISPR–Cas9-engineered T cells. Cell 186, 4567–4582 (2023).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Li, X. et al. Base editing with a Cpf1–cytidine deaminase fusion. Nat. Biotechnol. 36, 324–327 (2018).
Lei, L. et al. APOBEC3 induces mutations during repair of CRISPR–Cas9-generated DNA breaks. Nat. Struct. Mol. Biol. 25, 45–52 (2018).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Yang, L. & Chen, J. A tale of two moieties: rapidly evolving CRISPR/Cas-based genome editing. Trends Biochem. Sci. 45, 874–888 (2020).
Kim, J. S. & Chen, J. Base editing of organellar DNA with programmable deaminases. Nat. Rev. Mol. Cell Biol. 25, 34–45 (2024).
Karimian, A., Ahmadi, Y. & Yousefi, B. Multiple functions of p21 in cell cycle, apoptosis and transcriptional regulation after DNA damage. DNA Repair 42, 63–71 (2016).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Kim, D. et al. Genome-wide target specificities of CRISPR RNA-guided programmable deaminases. Nat. Biotechnol. 35, 475–480 (2017).
Wang, X. et al. Efficient base editing in methylated regions with a human APOBEC3A–Cas9 fusion. Nat. Biotechnol. 36, 946–949 (2018).
Mayuranathan, T. et al. Potent and uniform fetal hemoglobin induction via base editing. Nat. Genet. 55, 1210–1220 (2023).
Liao, J. et al. Therapeutic adenine base editing of human hematopoietic stem cells. Nat. Commun. 14, 207 (2023).
Cradick, T. J., Qiu, P., Lee, C. M., Fine, E. J. & Bao, G. COSMID: a web-based tool for identifying and validating CRISPR/Cas off-target sites. Mol. Ther. Nucleic Acids 3, e214 (2014).
Cromer, M. K. et al. Comparative analysis of CRISPR off-target discovery tools following ex vivo editing of CD34+ hematopoietic stem and progenitor cells. Mol. Ther. 31, 1074–1087 (2023).
Wang, L. et al. Enhanced base editing by co-expression of free uracil DNA glycosylase inhibitor. Cell Res. 27, 1289–1292 (2017).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Yang, B., Yang, L. & Chen, J. Development and application of base editors. CRISPR J. 2, 91–104 (2019).
Taher, A., Vichinsky, E., Musallam, K., Cappellini, M. D. & Viprakasit, V. in Guidelines for the Management of Non Transfusion Dependent Thalassaemia (NTDT) (ed. Weatherall, D.) (Thalassaemia International Federation, 2013).
Kirk, P. et al. International reproducibility of single breathhold T2* MR for cardiac and liver iron assessment among five thalassemia centers. J. Magn. Reson. Imaging 32, 315–319 (2010).
Sharma, A. et al. CRISPR–Cas9 editing of the HBG1 and HBG2 promoters to treat sickle cell disease. N. Engl. J. Med. 389, 820–832 (2023).
Frangoul, H. et al. Exagamglogene autotemcel for severe sickle cell disease. N. Engl. J. Med. 390, 1649–1662 (2024).
Fucharoen, S., Shimizu, K. & Fukumaki, Y. A novel C-T transition within the distal CCAAT motif of the Gγ-globin gene in the Japanese HPFH: implication of factor binding in elevated fetal globin expression. Nucleic Acids Res. 18, 5245–5253 (1990).
Oner, R., Kutlar, F., Gu, L. H. & Huisman, T. H. The Georgia type of nondeletional hereditary persistence of fetal hemoglobin has a C→T mutation at nucleotide-114 of the Aγ-globin gene. Blood 77, 1124–1125 (1991).
Motum, P. I., Deng, Z. M., Huong, L. & Trent, R. J. The Australian type of nondeletional Gγ-HPFH has a C→G substitution at nucleotide -114 of the Gγ gene. Br. J. Haematol. 86, 219–221 (1994).
Zertal-Zidani, S. et al. A novel C→A transversion within the distal CCAAT motif of the Gγ-globin gene in the Algerian Gγβ+-hereditary persistence of fetal hemoglobin. Hemoglobin 23, 159–169 (1999).
Ribeil, J. A. et al. Ineffective erythropoiesis in β-thalassemia. Sci. World J. 2013, 394295 (2013).
Cancellieri, S. et al. Human genetic diversity alters off-target outcomes of therapeutic gene editing. Nat. Genet. 55, 34–43 (2023).
Breda, L. et al. In vivo hematopoietic stem cell modification by mRNA delivery. Science 381, 436–443 (2023).
Wognum, B., Yuan, N., Lai, B. & Miller, C. L. Colony forming cell assays for human hematopoietic progenitor cells. Methods Mol. Biol. 946, 267–283 (2013).
Mathias, L. A. et al. Ineffective erythropoiesis in β-thalassemia major is due to apoptosis at the polychromatophilic normoblast stage. Exp. Hematol. 28, 1343–1353 (2000).
Arlet, J. B. et al. HSP70 sequestration by free α-globin promotes ineffective erythropoiesis in β-thalassaemia. Nature 514, 242–246 (2014).
Acknowledgements
This work is supported by the National Key Research and Development Program of China (2024YFC3405902 to L.Y., 2023YFC3403402 to J.C., 2024YFA1803301 to J.C. and 2023ZD0500501 to B.Y.), the National Natural Science Foundation of China (32530057 to J.C., 32371514 to J.C., 32430018 to L.Y. and 32070170 to B.Y.), the Agriculture Science and Technology Major Project (to J.C.), the Shanghai Municipal Science and Technology Commission (23JS1400300 and 23DX1900102 to L.Y., 23ZR1442500 to B.Y., and 23YF1407400 to X.-K.M.), the Program of Shanghai Academic/Technology Research Leader (23XD1422500 to J.C.) and the Program of Shanghai Rising-Star (24QB2707400 to L.W.). We thank staff at the Molecular and Cell Biology Core Facility, School of Life Science and Technology, ShanghaiTech University for their technical support; and K. Lan, Y. Zhang and P. Li at CorrectSequence Therapeutics for their technical support. This work was also supported in part by a Shanghai Municipal Education Commission (SMEC) grant to the Shanghai Frontiers Science Centre for Biomacromolecules and Precision Medicine at the ShanghaiTech University. J.C. is a SANS Exploration Scholar.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.C., Y. Lai, R.L., X.M. and L.Y. Investigation: Y. Lai, R.L., G.Y., L.S., Z.W. and X.Z. Statistical analysis: L.W. and X.-K.M. Methodology: L.W., X.-K.M., W.X., Y.H., Y.-L.G. and B.Y. Project administration: Y. Li and X.M. Access to all data: J.C. and L.Y. Writing original draft: J.C and L.W. Writing, reviewing and editing: J.C., L.W., L.Y., Y. Lai, R.L., X.-K.M., A.M., Y. Li and X.M. All authors agreed to submit the manuscript, read and approved the final draft and take full responsibility of its content.
Corresponding authors
Ethics declarations
Competing interests
J.C., L.Y. and B.Y. are scientific cofounders of CorrectSequence Therapeutics, a company that uses gene-editing technologies, and the members of its scientific advisory board. X.M., L.W. and Y. Li are employees of CorrectSequence Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks M. Kyle Cromer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Determination of indel frequencies, HbF levels, and F-cell percentages in healthy donor HSPCs and DNA damage signaling in patient HSPCs.
a, Frequencies of small indels in tBE-edited healthy donor HSPCs. The data are presented as the mean ± SD; n = 5 biologically independent CD34+ HSPC samples. b, HPLC data of HbF levels in Lyphochek Haemoglobin A2 Control and erythroid cells differentiated from tBE-edited healthy donor HSPCs. c, FACS analysis of F-cell percentages in erythroid cells differentiated from tBE-edited healthy donor HSPCs. d, Real-time qPCR analysis of CDKN1A (p21) transcript levels in primary patient–derived HSPCs following treatment with Cas9 nuclease or tBE using the clinical-scale manufacturing process. The data are presented as the mean ± SD; n = 2 biologically independent CD34+ patient HSPC samples; each sample was measured in four technical replicates.
Extended Data Fig. 2 Determination of the engraftment efficiency of tBE-edited CD34+ HSPCs from healthy donors.
a, FACS analysis of human cells in the bone marrow of mice engrafted with unedited or tBE-edited healthy donor CD34+ HSPCs. b, Representative flow cytometry staining and gating scheme for human cell engraftment in mouse bone marrow. Human B cells were gated as hCD3−/hCD19+. Human T cells were gated as hCD3+/hCD19−. Human myeloid cells were gated as hCD3−/hCD19−/hCD33+.
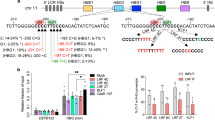
Extended Data Fig. 4 Comparison of tBE with other base editing approaches using HSPCs from two representative patients.
a, Sequences of sgRNA spacer regions covering the BCL11A binding motifs, along with base editing frequencies induced by different base-editing tools using the indicated sgRNAs. b, HPLC analysis of HbF levels and FACS quantification of F-cell percentages in erythroid cells differentiated in vitro from edited HSPCs. c, Comparison of off-target (OT) editing frequencies induced by the indicated base editors with the indicated sgRNAs at 145 potential off-target sites: 87 predicted by Cas-OFFinder, 36 identified by Digenome-seq, and 22 predicted by COSMID. For tBE and hA3A-BE3, data represent C-to-T/A/G conversion frequencies; for ABE8e, data represent mean A-to-G/C/T conversion frequencies. d, Comparison of OT editing frequencies induced by the indicated base editors with the indicated sgRNAs at 28 potential off-target sites. For hA3A (N57Q)-BE3, data represent C-to-T/A/G conversion frequencies; for ABE8e, data represent mean A-to-G/C/T conversion frequencies. e, Comparison of OT editing frequencies induced by the indicated editors with the indicated sgRNAs at 12 potential off-target sites. In b-e, the data are presented as the mean ± SD; n = 2.
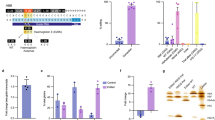
Extended Data Fig. 5 CESSCO assay and representative blood cell subtype flow cytometry analysis.
a, Schematic diagrams illustrate the co-expressing S. aureus and S. pyogenes Cas9 orthologs (CESSCO) assay, which expresses two Cas9 orthologs, to detect guide-RNA-independent off-target mutations induced by base editors at the R-loop region generated by nSaCas9. b, On-target editing frequencies induced by tBE and hA3A-BE3 in the presence of nSaCa9 and the indicated sgRNAs for S. aureus (Sa-sgRNAs). c, Off-target editing frequencies induced by tBE and hA3A-BE3 at the R-loop regions formed by nSaCas9 and the indicated Sa-sgRNAs. For b and c, the data are presented as the mean ± SD; n = 3 biologically independent 293 FT cells. d, Representative flow cytometry staining and gating strategy for patient blood cell subtypes. Granulocytes were gated as hCD45+ with high SSC. Monocytes were gated as hCD45+/hCD14+ with low SSC. B cells were gated as hCD45+/hCD3−/hCD19+. T cells were gated as hCD45+/hCD3+/hCD56−/hCD16−. NK cells were gated as hCD45+/hCD3−/hCD56+/hCD16+. Schematic in a adapted from ref. 5, Springer Nature Limited.
Extended Data Fig. 6 Manufacture and pharmacology data of CS-101, and summary of laboratory assessments in study patients.
a, Schematic of CS-101 manufacturing and infusion workflow. b, tBE-mediated base editing efficiency across 12 clinical-scale CS-101 batches. c, Cell viability following tBE-mediated base editing across 12 clinical-scale CS-101 batches. In b-c, the data are presented as the mean ± SD; n = 12 biologically independent CD34+ HSPCs. d, Base editing efficiency in engrafted cells from mouse bone marrow or peripheral blood at 16 weeks post-transplantation relative to input cells. The data are presented as the mean ± SD. For CS-101_Mock Cell engrafted mice, n = 8 biologically independent mice for bone marrow and peripheral blood cells. For CS-101_Edited Cell engrafted mice, n = 9 and n = 7 biologically independent mice for bone marrow and peripheral blood cells, respectively. e, Real-time qPCR analysis of γ-globin mRNA levels relative to β-like globin in engrafted cells from mouse bone marrow or peripheral blood at 16 weeks post-transplantation, compared with mock-treated cells. The data are presented as the mean ± SD. For CS-101_Mock Cell engrafted mice, n = 8 biologically independent mice for bone marrow and peripheral blood cells. For CS-101_Edited Cell engrafted mice, n = 9 and n = 8 biologically independent mice for bone marrow and peripheral blood cells, respectively. f, Neutrophil engraftment time stratified by spleen status. g, Total haemoglobin levels at each visit, overall and stratified by genotype, age, or sex. h, HbF levels at each visit, overall and stratified by genotype, age, or sex. i, Summary of erythropoiesis-associated laboratory parameters. In patients with β-thalassaemia, regular transfusion of normal donor red blood cells results in a substantial proportion of circulating red cells having normal volume and haemoglobin content, which keeps some base-line laboratory values within or close to the normal range.
Extended Data Fig. 7 Determination of HbF levels and F-cell percentages in one representative study patient and haemoglobin concentrations in all patients.
a, HbF levels in erythroid cells of one representative study patient measured by capillary electrophoresis. b, FACS analysis of the percentage of F-cells among the erythroid cells of one representative study patient. c. The expression levels of total Hb, HbF, HbA, and HbA2 over time in all five study patients.
Extended Data Fig. 8 Frequency of small indels in study patients and EMSA analysis of different editing outcomes.
a, Frequencies of small indels in CS-101 product. b, Frequencies of small indels in PBMCs over time in the study patients. c, Frequencies of small indel in bone marrow blood cells over time in the study patients. d, EMSA to analyse the binding of BCL11A to the top 10 mutated BCL11A binding motifs. Each experiment was independently repeated 3 times with similar results and representative data from one experiment are shown. The source image can be found in Supplementary Fig. 1.
Extended Data Fig. 9 Haemoglobin chain composition and iron overload parameters after CS-101 infusion.
a, RP-HPLC profiles of haemoglobin monomers in PBMCs and bone marrow cells from study patients at the indicated time points. γA represents γ-globin encoded by HBG1; γG represents γ-globin encoded by HBG2; α, β, and δ denote α-globin, β-globin, and δ-globin, respectively. Red arrows indicate individual peaks, and red dashed lines mark the start and end of each peak. b, Percentage-stacked histograms showing the distribution of haemoglobin monomers in peripheral blood and bone marrow cells from five patients over time. c, FACS analysis of the percentage of apoptotic cells in bone marrow erythroid cells from Patient 2. d, Change in serum ferritin level from baseline over time. e-f, Change in liver iron concentration and cardiac iron content from baseline, as assessed by T2*-weighted MRI.
Extended Data Fig. 10 Grade 3–4 adverse events and single-cell editing outcomes after CS-101 infusion.
a, Timeline of onset and resolution of all Grade 3 or 4 adverse events observed after CS-101 infusion. b, Proportion of single-cell–derived erythroid and granulocyte/monocyte colonies from patient bone marrow containing zero to four productive on-target edits at the BCL11A binding motifs in the HBG1/2 promoter at the indicated time points after infusion.
Supplementary information
Supplementary Information (download PDF )
This Supplementary Information file contains Supplementary Fig. 1, Supplementary Tables 1–8 and study protocol v.1.2.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lai, Y., Liu, R., Wang, L. et al. Clinical application of base editing for treating β-thalassaemia. Nature (2026). https://doi.org/10.1038/s41586-026-10342-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10342-9