Abstract
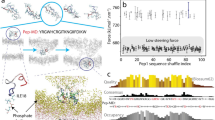
Peptide nanodiscs are promising anti-atherosclerosis therapeutics, drug delivery particles and structural biology tools. However, the lack of experimental methods for structural and dynamical characterization of these particles hinders their further development. Here, we integrated nuclear magnetic resonance (NMR), small-angle x-ray scattering, and small-angle neutron scattering experiments with molecular dynamics (MD) simulations to investigate the structure and dynamics of peptide nanodiscs stabilized by the apolipoprotein A-I mimetic peptide 22 A with therapeutic activity against atherosclerosis. This multi-technique approach takes advantage of combining average size and shape information from small-angle scattering, peptide site-specific information from NMR spectroscopy, and interpretative power of MD simulations. Our results reveal the intrinsic polydispersity in the size of peptide nanodiscs. Our consensus model suggests that 22 A peptides are predominantly in α-helical configuration with a disordered inter-helical orientation around the lipid matrix. The terminal regions of the peptides display greater flexibility relative to the peptide core and an enhanced C-terminal exposure to solvent, which could facilitate interaction with the enzyme LCAT. The methodological approach described in this paper paves the way for the design of more stable and effective therapeutic nanodiscs and for the characterization of other biomolecular aggregates that are beyond the scope of current structural biology techniques.
Similar content being viewed by others
Data availability
Molecular dynamics data and NMR spectra are available at https://zenodo.org/doi/10.5281/zenodo.18773356. All other relevant data are available upon request from the corresponding authors.
References
Osei-Hwedieh, D. O., Amar, M., Sviridov, D. & Remaley, A. T. Apolipoprotein mimetic peptides: Mechanisms of action as anti-atherogenic agents. Pharmacol. Therapeutics 130, 83–91 (2011).
Navab, M. et al. Peptide mimetics of apolipoproteins improve HDL function. J. Clin. Lipidol. 1, 142–147 (2007).
Patel, H. et al. Characterization of apolipoprotein A-I peptide phospholipid interaction and its effect on HDL nanodisc assembly. IJN 14, 3069–3086 (2019).
Anada, C., Ikeda, K., Nakao, H. & Nakano, M. Improvement of Thermal Stability of Amphipathic Peptide–Phospholipid Nanodiscs via Lateral Association of α-Helices by Disulfide Cross-Linking. Langmuir 38, 6977–6983 (2022).
Kingwell, B. A., Chapman, M. J., Kontush, A. & Miller, N. E. HDL-targeted therapies: progress, failures and future. Nat. Rev. Drug Discov. 13, 445–464 (2014).
Anantharamaiah, G.M. & Goldberg, D. Eds., Apolipoprotein Mimetics in the Management of Human Disease (Springer International Publishing, 2015).
Sviridov, D. & Remaley, A. T. High-density lipoprotein mimetics: promises and challenges. Biochemical J. 472, 249–259 (2015).
Kuai, R., Li, D., Chen, Y. E., Moon, J. J. & Schwendeman, A. High-Density Lipoproteins: Nature’s Multifunctional Nanoparticles. ACS Nano 10, 3015–3041 (2016).
Yuan, W. et al. Systematic evaluation of the effect of different apolipoprotein A-I mimetic peptides on the performance of synthetic high-density lipoproteins in vitro and in vivo. Nanomed.: Nanotechnol., Biol. Med. 48, 102646 (2023).
Midtgaard, S. R. et al. Self-assembling peptides form nanodiscs that stabilize membrane proteins. Soft Matter 10, 738–752 (2014).
Zhang, M. et al. Reconstitution of the Cyt b5 –CytP450 Complex in Nanodiscs for Structural Studies using NMR Spectroscopy. Angew. Chem. Int Ed. 55, 4497–4499 (2016).
Krishnarjuna, B. et al. Characterization of nanodisc-forming peptides for membrane protein studies. J. Colloid Interface Sci. 653, 1402–1414 (2024).
Krishnarjuna, B., Anantharamaiah, G. M. & Ramamoorthy, A. Peptide nanodiscs: Versatile platforms for membrane protein functional reconstitution and structural studies: A review. Int. J. Biol. Macromolecules 333, 148668 (2025).
Elzoghby, A. O. et al. Nanodiscs: Game changer nano-therapeutics and structural biology tools. Nano Today 53, 102026 (2023).
Hagn, F., Nasr, M. L. & Wagner, G. Assembly of phospholipid nanodiscs of controlled size for structural studies of membrane proteins by NMR. Nat. Protoc. 13, 79–98 (2018).
Günsel, U. & Hagn, F. Lipid Nanodiscs for High-Resolution NMR Studies of Membrane Proteins. Chem. Rev. 122, 9395–9421 (2022).
Bibow, S. et al. Solution structure of discoidal high-density lipoprotein particles with a shortened apolipoprotein A-I. Nat. Struct. Mol. Biol. 24, 187–193 (2017).
Bengtsen, T. et al. Structure and dynamics of a nanodisc by integrating NMR, SAXS and SANS experiments with molecular dynamics simulations. eLife 9, e56518 (2020).
Skar-Gislinge, N., Johansen, N. T., Høiberg-Nielsen, R. & Arleth, L. Comprehensive Study of the Self-Assembly of Phospholipid Nanodiscs: What Determines Their Shape and Stoichiometry? Langmuir 34, 12569–12582 (2018).
Xu, D. et al. Reconfigurable Peptide Analogs of Apolipoprotein A-I Reveal Tunable Features of Nanodisc Assembly. Langmuir 39, 1262–1276 (2023).
Salnikov, E. S., Anantharamaiah, G. M. & Bechinger, B. Supramolecular Organization of Apolipoprotein-A-I-Derived Peptides within Disc-like Arrangements. Biophysical J. 115, 467–477 (2018).
Pourmousa, M. & Pastor, R. W. Molecular dynamics simulations of lipid nanodiscs. Biochimica et. Biophysica Acta (BBA) - Biomembranes 1860, 2094–2107 (2018).
Islam, R. M. et al. Structural properties of apolipoprotein A-I mimetic peptides that promote ABCA1-dependent cholesterol efflux. Sci. Rep. 8, 2956 (2018).
Nencini, R. et al. Probing the dynamic landscape of peptides in molecular assemblies by synergized NMR experiments and MD simulations. Commun. Chem. 7, 28 (2024).
Sandelin, A. E. et al. Quality Evaluation Based Simulation Selection (QEBSS) for analysis of conformational ensembles and dynamics of multidomain proteins. Commun. Chem. 8, 241 (2025).
Dasseux, J.-L. et al. Apolipoprotein A-I agonist and their use to treat dyslipidemic disorders, (1999).
Li, D., Gordon, S., Schwendeman, A. & Remaley, A.T. “Apolipoprotein Mimetic Peptides for Stimulating Cholesterol Efflux” in Apolipoprotein Mimetics in the Management of Human Disease, G. M. Anantharamaiah, D. Goldberg, Eds. (Springer International Publishing, 2015), pp. 29–42.
Giorgi, L. et al. Mechanistic Insights into the Activation of Lecithin–Cholesterol Acyltransferase in Therapeutic Nanodiscs Composed of Apolipoprotein A-I Mimetic Peptides and Phospholipids. Mol. Pharmaceutics 19, 4135–4148 (2022).
Graziano, V., Miller, L. & Yang, L. Interpretation of solution scattering data from lipid nanodiscs. J. Appl Crystallogr 51, 157–166 (2018).
Skar-Gislinge, N. et al. Elliptical Structure of Phospholipid Bilayer Nanodiscs Encapsulated by Scaffold Proteins: Casting the Roles of the Lipids and the Protein. J. Am. Chem. Soc. 132, 13713–13722 (2010).
Cavanagh, J., Fairbrother, W.J., Palmer, A. G. III, Rance, M. & Skelton, N. J. Protein NMR Spectrocopy (Second Edition), Academic Press (2007).
Cooke, A. L. et al. A thumbwheel mechanism for APOA1 activation of LCAT activity in HDL[S]. J. Lipid Res. 59, 1244–1255 (2018).
Ollila, O. H. S., Heikkinen, H. A. & Iwaï, H. Rotational Dynamics of Proteins from Spin Relaxation Times and Molecular Dynamics Simulations. J. Phys. Chem. B 122, 6559–6569 (2018).
Kay, L. E., Torchia, D. A. & Bax, A. Backbone dynamics of proteins as studied by nitrogen-15 inverse detected heteronuclear NMR spectroscopy: application to staphylococcal nuclease. Biochemistry 28, 8972–8979 (1989).
Petrache, H. I., Dodd, S. W. & Brown, M. F. Area per Lipid and Acyl Length Distributions in Fluid Phosphatidylcholines Determined by 2H NMR Spectroscopy. Biophysical J. 79, 3172–3192 (2000).
Fawaz, M. V. et al. Phospholipid Component Defines Pharmacokinetic and Pharmacodynamic Properties of Synthetic High-Density Lipoproteins. J. Pharm. Exp. Ther. 372, 193–204 (2020).
Sibille, N. & Bernadó, P. Structural characterization of intrinsically disordered proteins by the combined use of NMR and SAXS. Biochemical Soc. Trans. 40, 955–962 (2012).
Chan-Yao-Chong, M., Durand, D. & Ha-Duong, T. Molecular Dynamics Simulations Combined with Nuclear Magnetic Resonance and/or Small-Angle X-ray Scattering Data for Characterizing Intrinsically Disordered Protein Conformational Ensembles. J. Chem. Inf. Model. 59, 1743–1758 (2019).
Reddy, T. & Rainey, J. K. Interpretation of biomolecular NMR spin relaxation parameters. Biochem. Cell Biol. 88, 131–142 (2010).
Debnath, A. & Schäfer, L. V. Structure and Dynamics of Phospholipid Nanodiscs from All-Atom and Coarse-Grained Simulations. J. Phys. Chem. B 119, 6991–7002 (2015).
Siuda, I. & Tieleman, D. P. Molecular Models of Nanodiscs. J. Chem. Theory Comput. 11, 4923–4932 (2015).
Shih, A. Y., Denisov, I. G., Phillips, J. C., Sligar, S. G. & Schulten, K. Molecular Dynamics Simulations of Discoidal Bilayers Assembled from Truncated Human Lipoproteins. Biophysical J. 88, 548–556 (2005).
Kjølbye, L. R. et al. General Protocol for Constructing Molecular Models of Nanodiscs. J. Chem. Inf. Model. 61, 2869–2883 (2021).
Mishra, V. K. et al. Association of a Model Class A (Apolipoprotein) Amphipathic α Helical Peptide with Lipid. J. Biol. Chem. 281, 6511–6519 (2006).
Sreerama, N. & Woody, R. W. Estimation of Protein Secondary Structure from Circular Dichroism Spectra: Comparison of CONTIN, SELCON, and CDSSTR Methods with an Expanded Reference Set. Anal. Biochem. 287, 252–260 (2000).
Zhu, G., Xia, Y., Nicholson, L. K. & Sze, K. H. Protein Dynamics Measurements by TROSY-Based NMR Experiments. J. Magn. Reson. 143, 423–426 (2000).
Delaglio, F. et al. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Skinner, S. P. et al. CcpNmr AnalysisAssign: a flexible platform for integrated NMR analysis. J. Biomol. NMR 66, 111–124 (2016).
Pernot, P., Round, A., Barrett, R. & De Maria Antolinos, A. al., Upgraded ESRF BM29 beamline for SAXS on macromolecules in solution. J. Synchrotron Rad. 20, 660–664 (2013).
Orthaber, D., Bergmann, A. & Glatter, O. SAXS experiments on absolute scale with Kratky systems using water as a secondary standard. J. Appl Crystallogr. 33, 218–225 (2000).
Förster, S., Apostol, L. & Bras, W. Scatter: software for the analysis of nano- and mesoscale small-angle scattering. J. Appl Crystallogr. 43, 639–646 (2010).
Manalastas-Cantos, K. et al. ATSAS 3.0: expanded functionality and new tools for small-angle scattering data analysis. J. Appl Crystallogr 54, 343–355 (2021).
Souza, P. C. T. et al. Martini 3: a general purpose force field for coarse-grained molecular dynamics. Nat. Methods 18, 382–388 (2021).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput Chem. 29, 1859–1865 (2008).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Dickson, C. J. et al. Lipid14: The Amber Lipid Force Field. J. Chem. Theory Comput. 10, 865–879 (2014).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Klauda, J. B. et al. Update of the CHARMM All-Atom Additive Force Field for Lipids: Validation on Six Lipid Types. J. Phys. Chem. B 114, 7830–7843 (2010).
Horn, H. W. et al. Development of an improved four-site water model for biomolecular simulations: TIP4P-Ew. J. Chem. Phys. 120, 9665–9678 (2004).
Berendsen, H. J. C., Van Der Spoel, D. & Van Drunen, R. GROMACS: A message-passing parallel molecular dynamics implementation. Computer Phys. Commun. 91, 43–56 (1995).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An N ⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Lu, C.-Y. & Vanden Bout, D. A. Effect of finite trajectory length on the correlation function analysis of single molecule data. J. Chem. Phys. 125, 124701 (2006).
Dayie, K. T., Wagner, G. & Lefèvre, J.-F. Theory and Practice of Nuclear Spin Relaxation in Proteins. Annu. Rev. Phys. Chem. 47, 243–282 (1996).
Svergun, D., Barberato, C. & Koch, M. H. J. CRYSOL – a Program to Evaluate X-ray Solution Scattering of Biological Macromolecules from Atomic Coordinates. J. Appl Crystallogr 28, 768–773 (1995).
Svergun, D. I. et al. Protein hydration in solution: Experimental observation by x-ray and neutron scattering. Proc. Natl. Acad. Sci. USA 95, 2267–2272 (1998).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Gowers, R. et al. MDAnalysis: A Python Package for the Rapid Analysis of Molecular Dynamics Simulations in (2016), pp. 98–105.
García De La Torre, J., Huertas, M. L. & Carrasco, B. Calculation of Hydrodynamic Properties of Globular Proteins from Their Atomic-Level Structure. Biophysical J. 78, 719–730 (2000).
Acknowledgements
We acknowledge the European Synchrotron Radiation Facility (ESRF) for provision of synchrotron radiation facilities under proposal number MX-2582 and we would like to thank Mark Tully for assistance and support in using beamline BM29. We acknowledge the Institut Laue-Langevin (ILL) for provision of neutron radiation facilities, and we would like to thank Anne Martel for assistance and support in using beamline D22. The facilities and expertise of the HiLIFE NMR unit at the University of Helsinki, a member of Instruct-ERIC Centre Finland, FINStruct, and Biocenter Finland are gratefully acknowledged. We acknowledge CSC – IT Center for Science for computational resources. This work was supported by The Academy of Finland [grant numbers 315596, 350636, 353815, 356568].
Author information
Authors and Affiliations
Contributions
S.I.V., R.N., and T.N. performed the NMR assignment. S.N performed and analyzed NMR and SAS experiments. A.N., R.N., and G.K performed and analyzed MD simulations. S.N, A.N., R.N., O.H.S.O., and A.K. wrote the manuscript. O.H.S.O. and A.K. conceptualized and supervised the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks Bankala Krishnarjuna, Evgeniy S. Salnikov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nouri, S., Niemelä, A., Nencini, R. et al. Exploring the structure and dynamics of peptide nanodiscs through a synergistic approach with NMR spectroscopy, SAS and MD simulations. Commun Chem (2026). https://doi.org/10.1038/s42004-026-02015-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-026-02015-5