Abstract
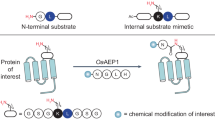
The ability to precisely modify proteins and peptides is fundamental to studying their function and creating new variants or topologies with improved properties. Recent studies have transformed the scope of transpeptidases as versatile tools for site-specific modification of proteins and peptides. The engineered asparaginyl ligase OaAEP1 is an ultrafast transpeptidase that stands out owing to its ability to efficiently catalyze a diverse range of modifications that extend well beyond its natural function to generate backbone cyclic peptides in plants. In this Protocol Extension, we describe a framework for the design and application of noncanonical reactions catalyzed by OaAEP1 that provide access to engineered products with customized terminal or side-chain modifications. The reactions proceed cleanly under mild, nondenaturing conditions and can be applied to a broad array of substrates produced by chemical synthesis or recombinant expression, including folded proteins and peptides. After preparing the required substrates and reagents (~5 d) and expressing the recombinant enzyme in Escherichia coli (~3 d), OaAEP1-catalyzed reactions can be carried out in a matter of minutes to hours. We describe methods for installing non-native C-terminal modifications, including by conjugating commercially available nonpeptidic amines (reactive handles, carbohydrates and so on) or ligating a reversed (retro) substrate mimetic that enables production of genetically inaccessible C-to-C fusions. We also describe procedures for OaAEP1-catalyzed side-chain modification of proteins and peptides, which can be applied to generate side-chain-to-tail macrocyclic products, to label a specific side-chain amine with a dye or other reporter tag, or to produce defined protein–cyclic peptide fusions.
Key points
-
This Protocol Extension describes a series of procedures for introducing different types of modifications into proteins or peptides using the asparaginyl ligase OaAEP1.
-
The reactions are highly versatile and enable late-stage modification of folded proteins and peptides, providing direct access to products with diverse C-terminal or side-chain modifications that are currently challenging to achieve with other enzymes or chemical approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
Data supporting the individual procedures detailed in this protocol are included in the current article and in the published articles that form the basis for the protocol.
References
Hoyt, E. A., Cal, P. M. S. D., Oliveira, B. L. & Bernardes, G. J. L. Contemporary approaches to site-selective protein modification. Nat. Rev. Chem. 3, 147–171 (2019).
Dumontet, C., Reichert, J. M., Senter, P. D., Lambert, J. M. & Beck, A. Antibody–drug conjugates come of age in oncology. Nat. Rev. Drug Discov. 22, 641–661 (2023).
Ebrahimi, S. B. & Samanta, D. Engineering protein-based therapeutics through structural and chemical design. Nat. Commun. 14, 2411 (2023).
Muttenthaler, M., King, G. F., Adams, D. J. & Alewood, P. E. Trends in peptide drug discovery. Nat. Rev. Drug Discov. 20, 309–325 (2021).
Rehm, F. B. H. et al. Asparaginyl ligases: new enzymes for the protein engineer’s toolbox. ChemBioChem 22, 2079–2086 (2021).
Harris, K. S. et al. Efficient backbone cyclization of linear peptides by a recombinant asparaginyl endopeptidase. Nat. Commun. 6, 10199 (2015).
Yap, K. et al. An environmentally sustainable biomimetic production of cyclic disulfide-rich peptides. Green Chem. 22, 5002–5016 (2020).
Yap, K. et al. Yeast-based bioproduction of disulfide-rich peptides and their cyclization via asparaginyl endopeptidases. Nat. Protoc. 16, 1740–1760 (2021).
Yang, R. et al. Engineering a catalytically efficient recombinant protein ligase. J. Am. Chem. Soc. 139, 5351–5358 (2017).
Rehm, F. B. H. et al. Site-specific sequential protein labeling catalyzed by a single recombinant ligase. J. Am. Chem. Soc. 141, 17388–17393 (2019).
Zhou, Y. et al. A chemoenzymatic approach to produce a cyclic analogue of the analgesic drug MVIIA (ziconotide). Angew. Chem. Int. Ed. 62, e202302812 (2023).
Le, T., Zhang, D., Martini, R. M., Biswas, S. & van der Donk, W. A. Use of a head-to-tail peptide cyclase to prepare hybrid RiPPs. Chem. Commun. 60, 6508–6511 (2024).
Du, J. et al. A bifunctional asparaginyl endopeptidase efficiently catalyzes both cleavage and cyclization of cyclic trypsin inhibitors. Nat. Commun. 11, 1575 (2020).
Li, C. Y. et al. Cystine knot peptides with tuneable activity and mechanism. Angew. Chem. Int. Ed. 61, e202200951 (2022).
Hemu, X. et al. Structural determinants for peptide-bond formation by asparaginyl ligases. Proc. Natl Acad. Sci. USA 116, 11737–11746 (2019).
Nguyen, G. K. T. et al. Butelase 1 is an Asx-specific ligase enabling peptide macrocyclization and synthesis. Nat. Chem. Biol. 10, 732–738 (2014).
Nguyen, G. K. T. et al. Butelase-mediated cyclization and ligation of peptides and proteins. Nat. Protoc. 11, 1977–1988 (2016).
Tang, T. M. S. et al. Use of an asparaginyl endopeptidase for chemoenzymatic peptide and protein labeling. Chem. Sci. 11, 5881–5888 (2020).
Rehm, F. B. H., Tyler, T. J., Yap, K., Durek, T. & Craik, D. J. Improved asparaginyl-ligase-catalyzed transpeptidation via selective nucleophile quenching. Angew. Chem. Int. Ed. 60, 4004–4008 (2021).
Chen, J. et al. Improvement of transpeptidative protein ligation via selective nucleophile quenching using 2-cyano-6-hydroxybenzothiazole. Org. Lett. 27, 3476–3481 (2025).
Xia, Y. et al. A cascade enzymatic reaction scheme for irreversible transpeptidative protein ligation. J. Am. Chem. Soc. 145, 6838–6844 (2023).
Nguyen, G. K. T., Cao, Y., Wang, W., Liu, C. F. & Tam, J. P. Site-specific N-terminal labeling of peptides and proteins using butelase 1 and thiodepsipeptide. Angew. Chem. Int. Ed. 54, 15694–15698 (2015).
Rehm, F. B. H., Tyler, T. J., de Veer, S. J., Craik, D. J. & Durek, T. Enzymatic C-to-C protein ligation. Angew. Chem. Int. Ed. 61, e202116672 (2022).
de Veer, S. J., Zhou, Y., Durek, T., Craik, D. J. & Rehm, F. B. H. Tertiary amide bond formation by an engineered asparaginyl ligase. Chem. Sci. 15, 5248–5255 (2024).
Rehm, F. B. H. et al. Enzymatic C-terminal protein engineering with amines. J. Am. Chem. Soc. 143, 19498–19504 (2021).
Rehm, F. B. H. et al. Repurposing a plant peptide cyclase for targeted lysine acylation. Nat. Chem. 16, 1481–1489 (2024).
de Veer, S. J., Craik, D. J. & Rehm, F. B. H. Highly efficient transpeptidase-catalyzed isopeptide ligation. J. Am. Chem. Soc. 147, 557–565 (2025).
Zhou, Y. et al. Asparaginyl ligases with engineered substrate specificity for controlled, sequential transpeptidation reactions. J. Am. Chem. Soc. 147, 21824–21831 (2025).
Hemu, X. et al. Turning an asparaginyl endopeptidase into a peptide ligase. ACS Catal 10, 8825–8834 (2020).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Boutureira, O. & Bernardes, G. J. Advances in chemical protein modification. Chem. Rev. 115, 2174–2195 (2015).
Glasgow, J. E., Salit, M. L. & Cochran, J. R. In vivo site-specific protein tagging with diverse amines using an engineered sortase variant. J. Am. Chem. Soc. 138, 7496–7499 (2016).
Weikart, N. D. & Mootz, H. D. Generation of site-specific and enzymatically stable conjugates of recombinant proteins with ubiquitin-like modifiers by the Cu(I)-catalyzed azide-alkyne cycloaddition. ChemBioChem 11, 774–777 (2010).
Islam, M. D., Islam, M. M. & Kuroda, Y. Ni2+-induced selective precipitation of His-tagged recombinant proteins shortens purification time while maintaining high yield. J. Biotechnol. 399, 38–46 (2025).
Li, H. et al. Peptidyl asparaginyl ligase-mediated orthogonal ligation at non-Asx peptide bonds for D-peptide cyclization and antibody dual labeling. J. Am. Chem. Soc. 148, 1989–1997 (2026).
Luo, J. et al. Capturing acyl-enzyme intermediates with genetically encoded 2,3-diaminopropionic acid for hydrolase substrate identification. Nat. Protoc. 19, 2967–2999 (2024).
Zhou, Y., de Veer, S. J., Durek, T. & Craik, D. J. AEP-mediated backbone cyclization of cyclotides—a chemoenzymatic method for cyclotide production. Methods Enzymol. 723, 71–90 (2025).
Guimaraes, C. P. et al. Site-specific C-terminal and internal loop labeling of proteins using sortase-mediated reactions. Nat. Protoc. 8, 1787–1799 (2013).
Cheneval, O. et al. Fmoc-based synthesis of disulfide-rich cyclic peptides. J. Org. Chem. 79, 5538–5544 (2014).
Flood, D. T. et al. Leveraging the Knorr pyrazole synthesis for the facile generation of thioester surrogates for use in native chemical ligation. Angew. Chem. Int. Ed. 57, 11634–11639 (2018).
Acknowledgements
Work in our laboratory on asparaginyl ligases is supported by a grant from the Australian Research Council (ARC, DP200101299) and by access to the facilities of the ARC Centre of Excellence for Innovations in Peptide and Protein Science (CE200100012). D.J.C. and F.B.H.R. are supported by National Health & Medical Research Council Investigator Grants (GNT2009564 to D.J.C. and GNT2018461 to F.B.H.R.).
Author information
Authors and Affiliations
Contributions
F.B.H.R. conceptualized the work and led the individual studies detailed in the protocol. S.J.d.V., Y.Z., F.B.H.R. and T.D. performed experiments and analyzed data. T.D. and D.J.C. supervised the work. The manuscript was written by S.J.d.V. and Y.Z. with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Yi-Ming Li, Yan-Mei Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Key references
de Veer, S. J. et al. J. Am. Chem. Soc. 147, 557–565 (2025): https://doi.org/10.1021/jacs.4c11964
Rehm, F. B. H. et al. Nat. Chem. 16, 1481–1489 (2024): https://doi.org/10.1038/s41557-024-01520-1
Rehm, F. B. H. et al. Angew. Chem. Int. Ed. 61, e202116672 (2022): https://doi.org/10.1002/anie.202116672
Rehm, F. B. H. et al. J. Am. Chem. Soc. 143, 19498–19504 (2021): https://doi.org/10.1021/jacs.1c08976
Yap, K. et al. Nat. Protoc. 16, 1740–1760 (2021): https://doi.org/10.1038/s41596-020-00483-0
This protocol is an extension to: Nat. Protoc. 16, 1740–1760 (2021): https://doi.org/10.1038/s41596-020-00483-0
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods and Figs. 1 and 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
de Veer, S.J., Zhou, Y., Rehm, F.B.H. et al. Site-specific protein and peptide modification by redeploying an asparaginyl ligase for noncanonical reactions. Nat Protoc (2026). https://doi.org/10.1038/s41596-026-01348-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41596-026-01348-8